what is this i don’t even
flat-mcmc is a Haskell library for painless, efficient, general-purpose sampling from continuous distributions.
Sampling is commonly used in Bayesian statistics/machine learning, physics, and finance to approximate difficult integrals or estimate model parameters. Haskell is an advanced functional language emphasizing abstraction, performance, multicore support, and security.
tell me more
Consider using a Gaussian likelihood model for some data. A conjugate prior yields a posterior that, in parameter space, looks like this:
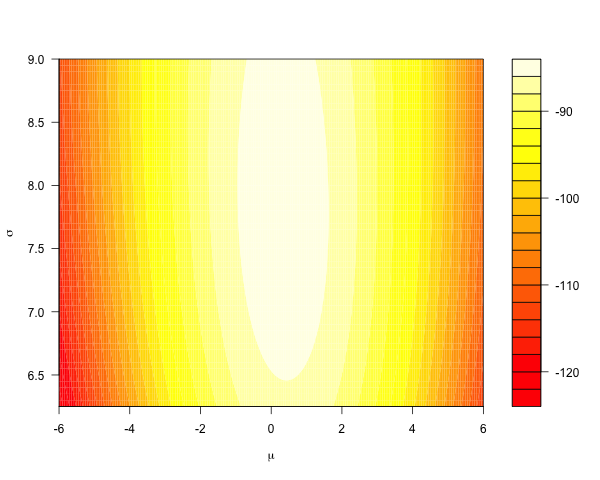
The Metropolis-Hastings algorithm is the ‘go-to’ general-purpose sampler, proposing global moves over both parameters simultaneously:
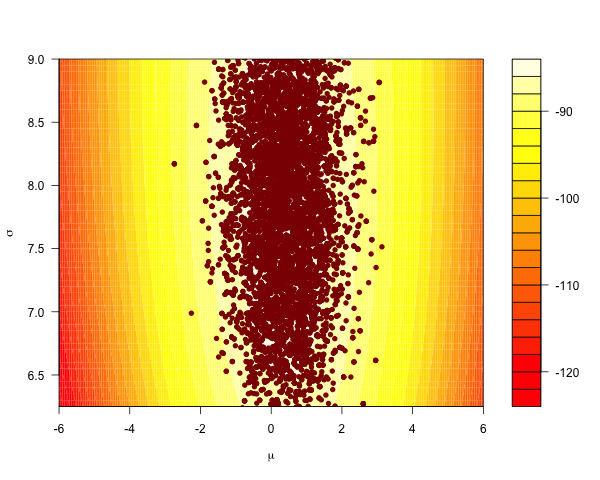
Distributions like this are easy to sample from efficiently. The case is different for anisotropic distributions that are ‘skewed’ or ‘stretched’. They look funny; narrow and correlated, like the Rosenbrock density on the plane:
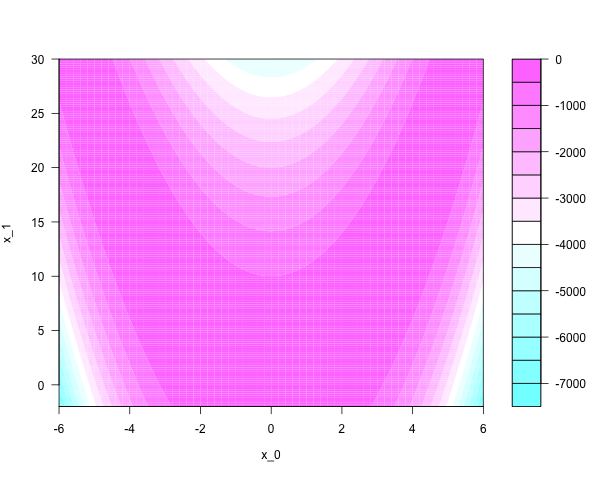
With some cleverness, the Rosenbrock density can be sampled independently. One thousand independent samples look like this:
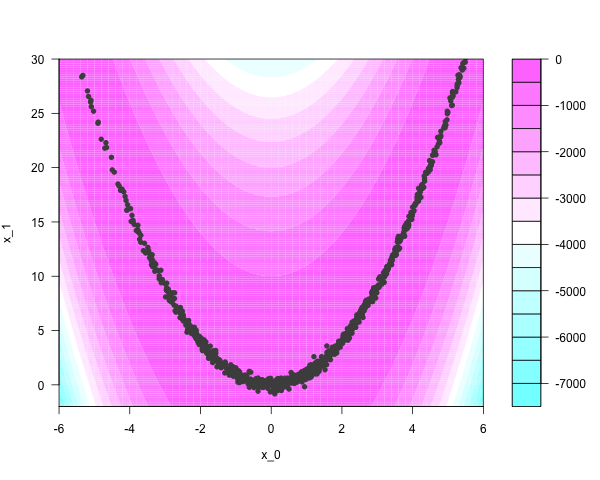
Conventional Markov chain samplers have trouble moving around narrow regions of the parameter space. Take fifty thousand iterations of a Metropolis-Hastings sampler, using naive Gaussian ‘bubble’ proposals:
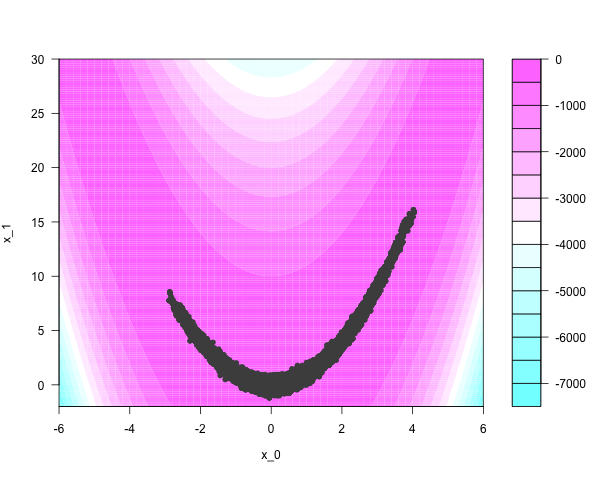
Tailoring good proposals requires local knowledge of the target manifold, which can be particularly unpleasant to incorporate in high dimensions.
Hamiltonian Monte Carlo (HMC) immediately finds regions of appreciable density and moves quickly through the parameter space. Three thousand iterations, at 20 discretizing steps per iteration, yields this:
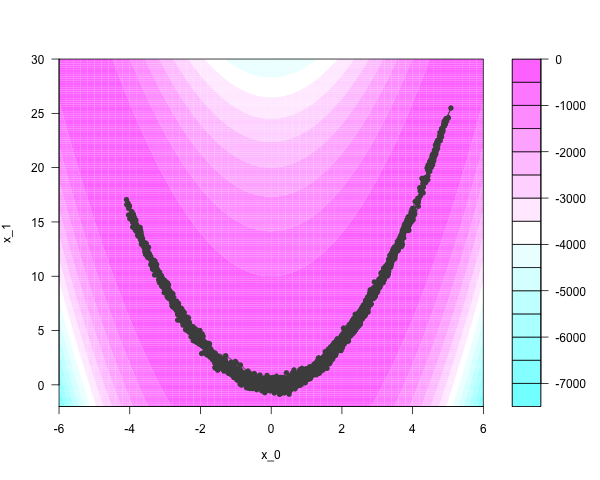
But HMC’s performance is very sensitive to the settings of two tuning parameters, and the algorithm usually requires preliminary calibration runs to get them right. Running the intermediate discretizing steps can also be time-consuming.
Another method - also quite quick, and requiring no tuning at all - involves ensemble samplers that are invariant to affine transformations of space. In essence, they ‘flatten’ or ‘unstretch’ the target’s parameter space, allowing many particles to explore the distribution locally. The equivalent work of the Metropolis-Hastings sampler yields something like this:
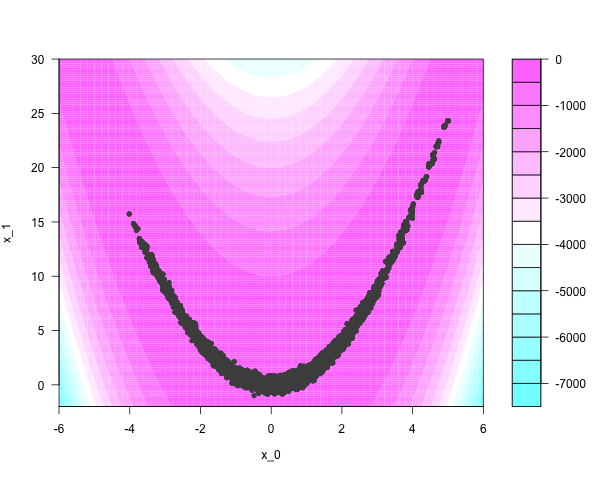
Good implementations of these algorithms exist. Haskell yields some perks; samplers can be compiled, and it is trivial to incorporate nested parallelism for specialized performance if compiled to GHC’s threaded runtime. flat-mcmc supports parallel evaluation of the target function via any of Haskell’s available methods:
and additionally, the ensemble’s particles are perturbed in parallel on each iteration of the Markov chain.
but does it blend
For smaller problems, the overhead of scheduling parallel computations will outweigh any gains made from using multiple cores. If the target function is fairly simple, or if the Markov ensemble contains relatively few particles, then it’s better to compile without the threaded runtime. Sequential performance is quite good:
Parallelism gains will manifest on problems with more expensive likelihoods, or when the Markov ensemble contains relatively many particles. Consider a 100-dimensional discretized stochastic PDE, expressed by the following recursive worker/wrapper function:
Run the chain for a million iterations, printing every 1000th to stdout; without the threaded runtime, the sequential sampler takes on the order of an hour:
But using four cores shaves about twenty minutes off:
join the club
Pull requests and bug reports welcome. If you find this work useful in your research, please cite it as
- Tobin, J. (2012) flat-mcmc: A library for efficient, general purpose sampling. jtobin.github.com/flat-mcmc
flat-mcmc is written and maintained by Jared Tobin. Page generated with Hakyll.